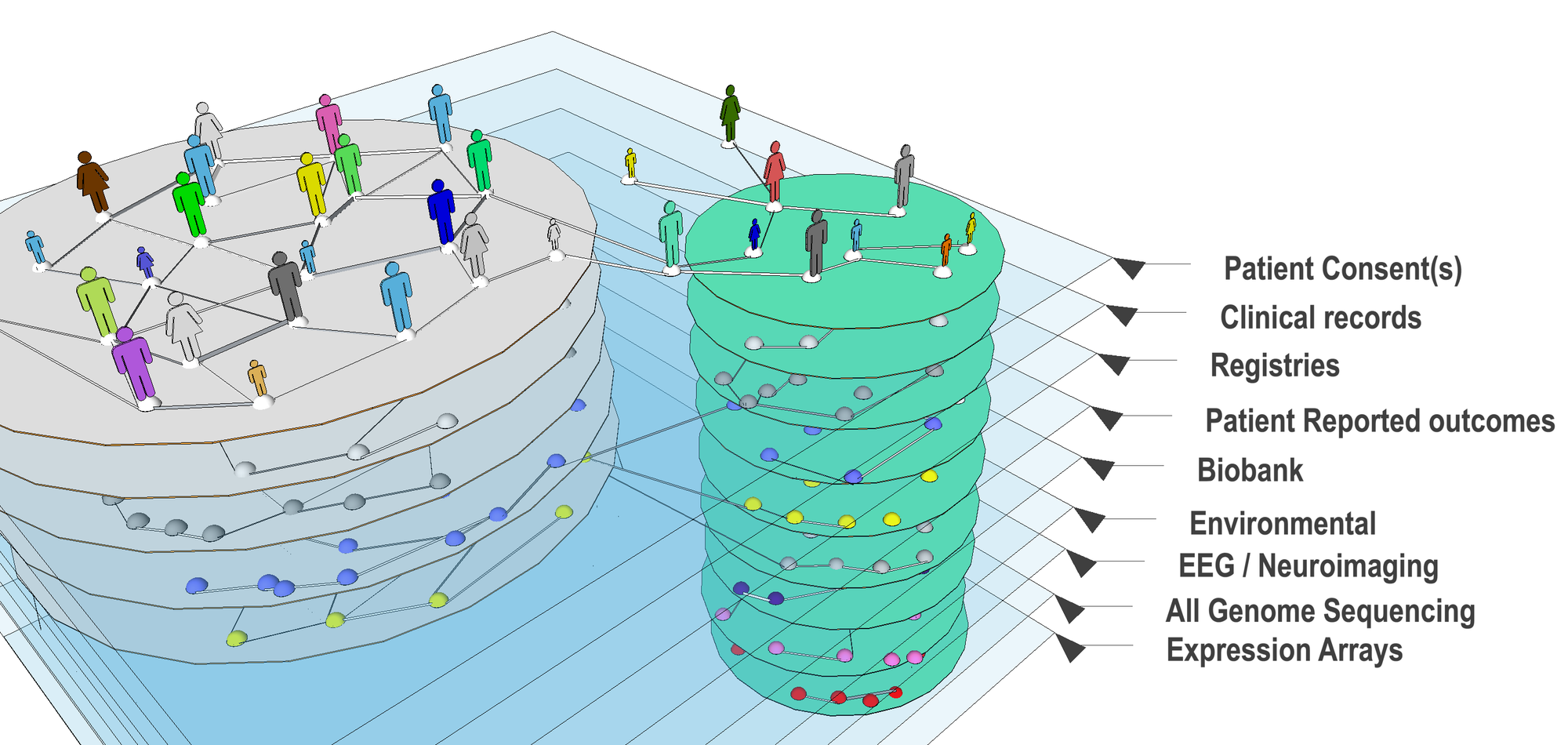
The ability to acquire and then reason over an entire spectrum of data types ranging from molecular and genomic all the way to clinical, epidemiological, environmental and social is now seen as central to improving patient care and advancing biomedical science.
This vision has led CHIP faculty to pursue interests in the methods, approaches, and tools of biomedical informatics and to an understanding that, increasingly, biomedical research and medical practice involve knowledge management and engineering and data-to-knowledge processes that can only be performed at scale by embracing the full spectrum of available data.